Highlights
Cell adhesions: the structures that hold life together
How biomolecular adhesions can adapt to mechanical load
Mechanical loading generally weakens adhesive structures and eventually leads to their rupture. However, biological systems can adapt to loads by strengthening adhesions, which is essential for maintaining the integrity of tissue and whole organisms. We have discovered a generic mechanism that can allow adhesion systems to harness applied loads for self-stabilization through growth. The mechanism is based on conformation changes of adhesion molecules that are dynamically exchanged with a reservoir. Load on the adhesion drives the occupation of some molecular states out of equilibrium, which, for thermodynamic reasons, leads to association of further molecules with the system. The pinciple of self-stabilization can be realized in many ways in complex adhesion-state networks and may contribute to adaptation of cellular focal adhesions involving the adaptor proteins talin and vinculin. Read more in our article published in Nature Communications.
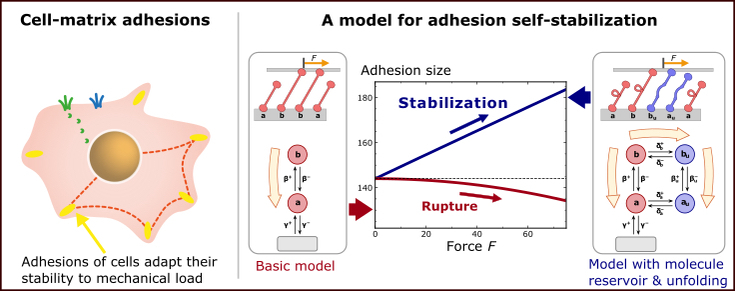
(Left) Sketch of a mammalian cell adhering to a planar substrate. Cell adhesions are micrometer-sized protein complexes that form a mechanical connection between the cell and its environment. Remarkably, cell-matrix adhesions can grow under force, thus adapting their stability to the applied mechanical load. (Right) We suggest a generic physical mechanism how load-dependent growth of adhesion sites can result from the combination of a conformation change of adhesion molecules and their exchange with a reservoir in the surrounding.
Stochastic micromechanics of bacteria
Bacteria "switch to a stronger gear" when forming a group
Bacterial migration, aggregation, and even host infection depend on the forces that bacteria exert on surfaces. Therefore, bacterial cell-surface forces play a prominent role in diseases, bacterial pollution, and biofouling. In spite of this importance, measuring forces between bacteria and surfaces has always been challenging. Recently, we succeeded in measuring these forces by using Myxococcus xanthus as a model organism. M. xanthus migrates on surfaces by employing two distinct molecular machineries, leading to two migration modes termed twitching and gliding. We were able to show that migration with these two modes produces very distinct surface forces. However, the two migration modes are similar in one surprising aspect, namely that forces from individual cells are amplified when the cells migrate in groups. During collective migration of M. xanthus, bacteria stick to each other while every single cell produces more force than it would individually. Thus, bacteria "switch to a stronger gear'' when they come together in a group. Read more in an our article that appeared in PNAS.
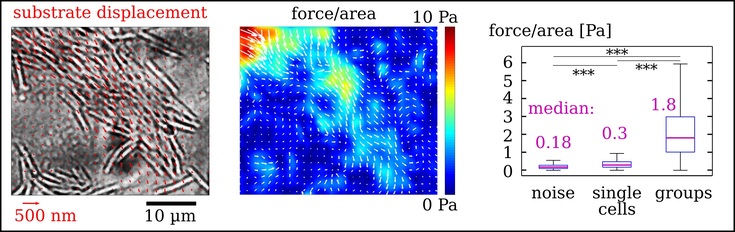
Traction force microscopy (TFM) measurements of substrate forces below a group of gliding myxobacteria. (Left) The bacteria deform an elastic substrate below them. (Middle) The substrate deformation allows to calculate force per area. (Right) In groups, bacteria produce much higher forces per area.
Theory of cellular sensing
How membrane channels can detect mechanical force
Sensing force is essential for life because this ability allows the survival of cells and organisms in changing environments. Force sensing by cells is often based on the micromechanics of membrane channels that open in response to forces. When opened, these channels allow the passage of signaling molecules, e.g., calcium ions, thus converting the mechanical stimulus to a chemical signal. This process is particularly well-studied in touch-sensing neurons and osmosensing neurons. In both cases, the membrane channels are attached to protein tethers that transmit forces directly to the channel. Recently, we proposed a model to explain how channel opening can be affected by the interplay of forces at a tether, tension in the cell membrane, and channel deformations. A central finding is that minute conical channel deformation under force leads to significant energy release during opening. This energy release may allow the direct detection of forces by tethered channels. Intriguingly, the sensitivity of force detection is amplified when more than one channel is present in the membrane. Channel-channel interactions via membrane deformations strongly affect the energetics of channel opening, which allows collective signal amplification. Read more in a paper by Benedikt Sabass and Howard Stone that appeared in Physical Review Letters.
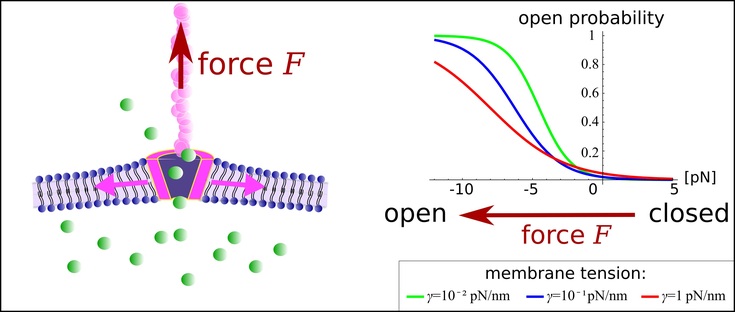
Role of the membrane for mechanosensing of tethered channels. (Left) Sketch of a channel in a membrane. The protein tether applies a force F on the channel. (Right) Our model predicts that certain channel molecules, e.g., TREK-1, respond to force at tethers by opening a pore, thus allowing conversion of a mechanical singnal to a chemical response.
Nonequilibrium thermodynamics of complex systems
Topology and Onsager symmetry: how linear networks become exciting
Oscillator networks are omnipresent; they occur for example in electric devices, in mechanical systems, and in biochemistry. However, purely linear networks have limited capabilities. Diode-like, monodirectional transport is usually impossible. Furthermore, maximum power cannot be transmitted with an efficiency greater than 1/2. Therefore, engineered networks often contain active or non-linear elements such as transistors. As a possible alternative, we investigate purely linear networks containing Lorentz-force-like couplings that break time-reversal symmetry. We found that these networks allow the construction of linear diodes with frequency-independent isolation properties. Moreover, the efficiency at maximum power can approach unity. These surprising properties require a combination of network loops with magnetic time-reversal symmetry breaking. Read more in an article by Benedikt Sabass that appeared in Europhysics Letters.
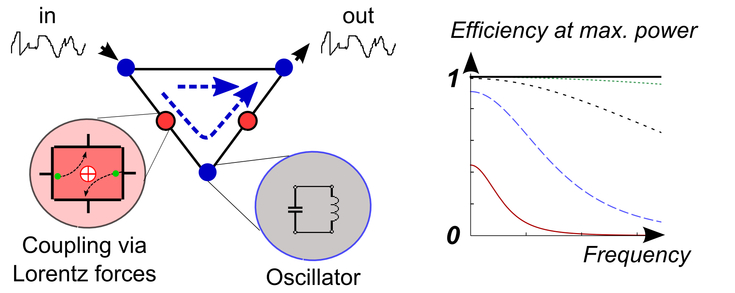
Sketch of a ''linear diode'' allowing perfect directional transport and efficiency at maximum power beyond 1/2.

